Call 800-462-3417 or email 2d@kendricklabs.com for a price quote. Download a catalog (PDF) or contact us to receive a printed catalog by mail.
2D
2D ELECTROPHORESIS
MAILING KIT (MK-1).
This kit contains ten 1 ml tubes of Urea Sample Buffer, ten 1 ml tubes of SDS Boiling Buffer, seventeen 1 ml tubes of SDS Boiling Buffer minus BME, seventeen 1 ml tubes of Osmotic Lysis Buffer, seventeen 100 μl tubes of 10X Nuclease Stock, four 100 μl tubes of Protease Inhibitor Stock Solution, and three 100 μl tubes of Phosphatase Inhibitor Stock Solutions I and II. The compositions of these solutions are provided in Suggestions for Sample Preparation. Along with the buffers and solutions mentioned, mailing instructions are included in the shipment.
Price:$418 includes shipping on dry ice.
2D ELECTROPHORESIS BASIC SERVICE (2D-ES-1).
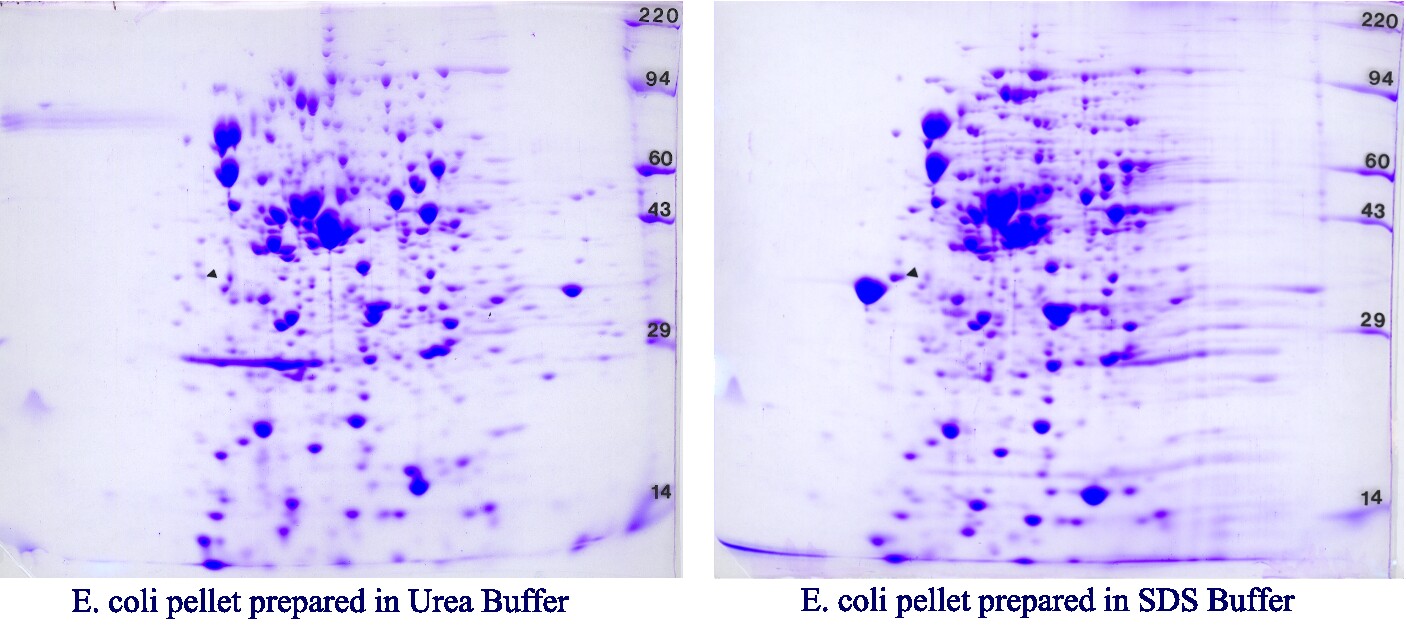
Our standard gel size is perfect for immunoprecipitations, subcellular fractionations, most bacterial preparations and many proteomics applications. One Coomassie blue-stained 2D gel of dried dimensions 15 cm wide by 13 cm long is returned for each sample along with a pH gradient plot and a method description sheet suitable for reports or publications. The example below is from an Escherichia coli sample prepared in our Urea sample buffer (left) and SDS boiling buffer (right) run identically on a pH 4-8 IEF gradient and 10% acrylamide slab gels. The advantage of SDS buffer is ease of sample preparation. Molecular weight standards (220, 94, 60, 43, 29, and 14 kDa) appear as bands on the right side of the gel labeled with press-on numbers. One isoelectric point marker (tropomyosin), added to each sample as an internal standard, is marked with a black arrow. Our standard IEF tube gel contains 2% pH 3-10 Isodalt or 2% pH 4-8 mix, both from Serva. Gels are transparency dried (STP-8) unless otherwise specified. See 2D-ES-5 for our large format option.
Price:2D-ES-1: 1 sample, $302; 2D-ES-1A: 2-3 samples, $278 each;
2D-ES-1B: 4-8 samples, $266 each; 2D-ES-1C: 9-20 samples, $249 each;
2D-ES-1D: 21+ samples, $237 each. 2D-Dup: duplicate gel, $142 each
CUSTOM PH GRADIENTS
Available (in addition to our standard pH 3-10 Isodalt gradient) using ampholines of pH 3.5-10, 2.5-5, and 7-9. For basic proteins with pI’s greater than 9, non-equilibrium pH gradient electrophoresis is used (NEPHGE, O’Farrell, P. et. al.Cell12: 1133, 1977). Buffers and electrodes are reversed and the pH gradient is created using pH 8-10.5 ampholines. Indicate which custom pH gradient you need on the back of the sample ID form. (No extra charge)
2D ELECTROPHORESIS WITH PEPTIDE SLAB GELS (2D-ES-2).
The 16.5% peptide slab gels of Schagger and von Jagow (Anal. Biochem. 166: 368, 1987) are useful to resolve low MW proteins (3 – 14 kDa). In general it is better to load low MW proteins heavily (10 μg/spot). Note that ampholines may interfere in the MW range 2-6 kDa.
Price: Add $49 per gel.
SEMI-NATIVE 2D ELECTROPHORESIS (2D-ES-3).
For semi-native 2D, denaturing agents urea, SDS, NP-40 and sulfhydral reagents are omitted from the sample buffer and from the second dimension slab gel. However, 9M urea is a structural component of the IEF tube gel and must be retained during isoelectric focusing. We don’t recommend this. Proteins tend to clump under semi-native conditions; some are insoluble and must be centrifuged off. However, in some cases this 2D variation is useful.
Price: Same as 2D-ES-1
2D ELECTROPHORESIS WITHOUT SULFHYDRAL REAGENT (2D-ES-4).
Same as 2D-ES-1 except 2-mercaptoethanol and dithiothreitol are omitted from sample buffer, internal standards, and all 2-D solutions.
Price: Same as 2D-ES-1
LARGE FORMAT 2D ELECTROPHORESIS (2D-ES-5).
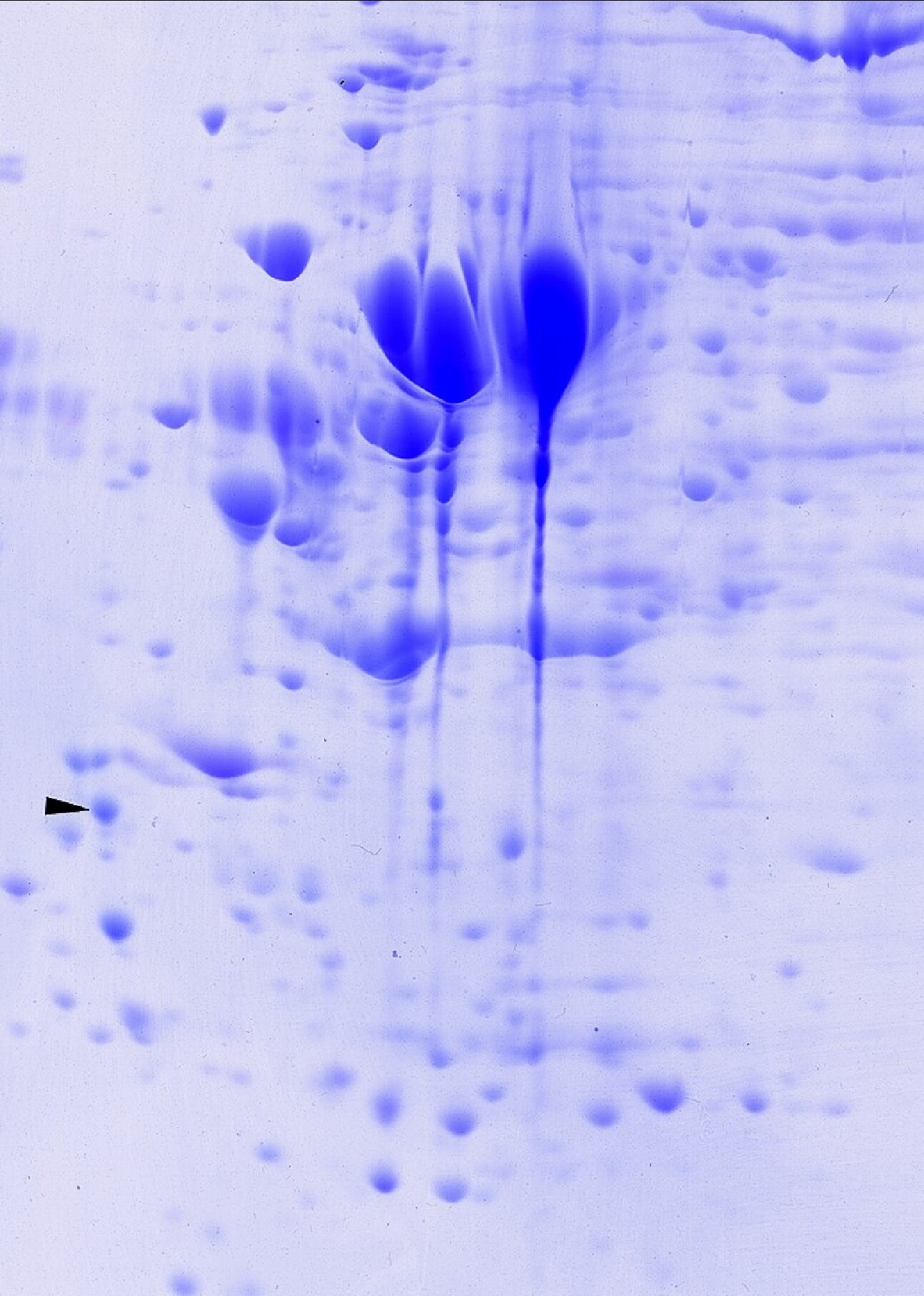
The larger size gives better resolution for proteomics applications; heavier loads may be applied to obtain more material for mass spectrometry. One Coomassie blue-stained 2D gel of dried dimensions 20 cm wide by 22 cm long is returned for each sample along with a pH gradient plot and a method description sheet. Our standard IEF tube gel contains 2% pH 3-10 Isodalt from Serva or 2% 4-8 ampholine mix. Molecular weight standards (220, 94, 60, 43, 29, and 14 kDa) appear as bands on the right side of the gel labeled with press-on numbers. One IEF marker (tropomyosin), added to each sample as an internal standard, is marked with a black arrow. Gels are transparency dried unless otherwise specified (STP-8) is standard. See above for custom pH gradients. Typical protein loads are 100 (silver staining) or 500 μg (Coomassie blue) in 25-150 μl as shown in images below.
Price: 2D-ES-5: 1 sample $443; 2D-ES-5A:2-3 samples $399 each;
2D-ES-5B: 4-12 samples $377 each; 2D-ES-5C: 13-200 samples $361 each;
LF-Dup: (Large Format 2D duplicate gels): $183 each.
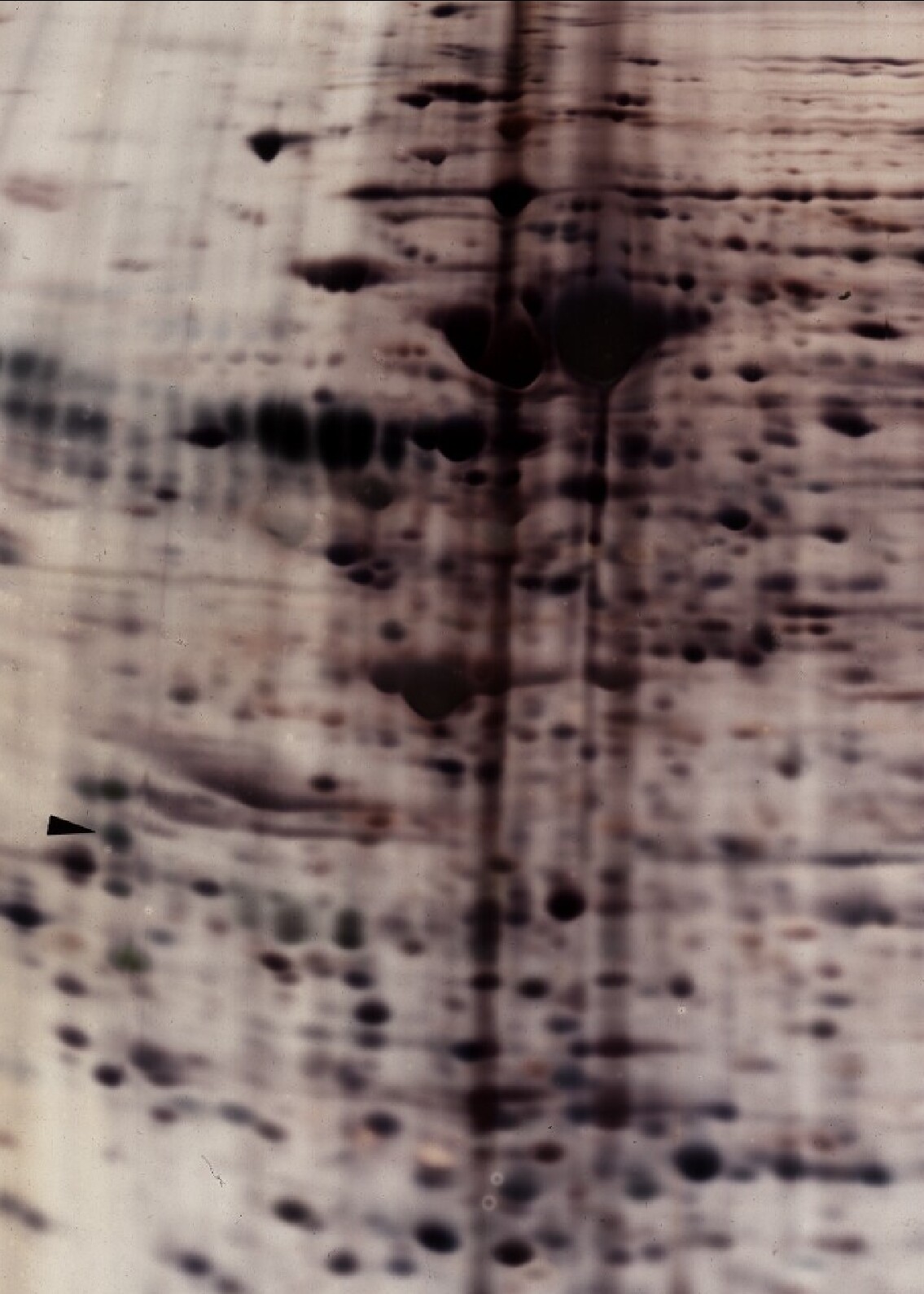


Large Format 2D Gel Examples, Silver versus Coomassie Staining.
A side by side comparison of 2D gels loaded with same sample and stained with Coomassie blue (left, 600 µg sample load) and with silver (right, 100 µg sample load). The silver stain is at least 10x more sensitive than Coomassie blue stain even though less is loaded.
CARBAMYLATED CARBONIC ANHYDRASE pI MARKERS (2D-ES-7).
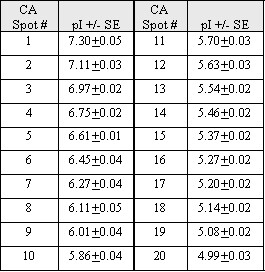
This protein was purchased from Sigma-Aldrich, progressively carbamylated on lysine residues by heating in urea buffer for various times, and calibrated for pI. Each spot in the beads-on-a-string pattern differs from the next by a single charge; measured pI values are shown in the table. A pH gradient standard curve may be determined from the train of charge isomers added as an internal standard for calculation of pIs for unknown proteins. A list of the isomers and their respective pI values are shown in the table below. Note that these pIs are not absolute values but are for conditions of 9 M urea and 22oC. The arrow marks an internal standard protein, tropomyosin, MW 33,000, pI 5.2. The right most spot in the isomer pattern on the gel is CA spot #1 in the table with a pI of 7.3.
Price: Add $35 per gel.

Comparisons
2D GEL COMPARISONS AND GEL SCANNING
MANUAL COMPARISONS OF 2D PATTERNS FOR DIFFERENCES (2D-ES-9).
We find that HBC (Human Brain Comparison) is a reliable method for finding changes in 2D patterns. In this method two experienced analysts independently compare stained gel pairs on a light box for differences. Results are presented as color coded outlines of spots on transparent overlay covering the gels. Free duplicate gels are run to confirm differences for samples scheduled for manual comparisons.
Price:2D-ES-9SF: $305 per Standard format pair compared
2D-ES-9LF: $355 per Large format pair compared
COMPUTERIZED COMPARISONS OF 2D GEL PATTERNS FOR DIFFERENCES (2D-ES-10).
This analysis includes scanning with a calibrated densitometer and computerized analysis with SameSpots software v4.5 (2010, TotalLab, Ltd.). Requires prior 2D electrophoresis but free duplicate gels are run and analyzed to confirm differences for samples scheduled for computerized comparisons. Generally 400-800 spots are quantified per Standard Format gel (13×15 cm); 800-2000 spots are quantified per Large Format gel (20×22 cm). Spot density values are expressed as spot percentages (individual spot density as a percentage of total density in all spots analyzed) to normalize for differences in sample loading or staining. Values for corresponding spots from duplicate gels are averaged.
Results are presented in a complete report including a summary table showing spot number, pI, MW, ratios (fold difference) and p values as a second measure of difference. The final report also includes figures of images showing numbering, montage images for every differing protein spot, methods and pH gradient plot. A CD containing data and image files is included in the package. Note that our reports focus on the protein differences between samples. Picking protein spots later for identification by mass spectrometry is straightforward. Just send a list of spot numbers and we’ll take it from there. Report Example
Price: 2D-ES-10: Standard format duplicate gels analyzed and averaged per sample. Per comparison, $798
2D-ES-10LF: Large format duplicate gels analyzed and averaged per sample. Per comparison, $860
Lic-1: Image licensing fee: Add $129/image
QUANTIFICATION OF PROTEINS RESOLVED BY 2-D ELECTROPHORESIS (2D-ES-12).
For clients knowing which proteins need to be quantified. Scanning (2D-ES-13) is included. Data is reported in integrated spot density units or as a percentage of total image density. Spots are not matched from gel to gel.
Price: 2D-ES-12: 1-12 spots, $162;
2D-ES-12A 13-50 spots, $278; 2D-ES-12B: 51-200 spots, $333
Lic-1: image licensing fee: Add $129/image
LASER DENSITOMETRY (2D-ES-13).
Stained gels and x-ray films are digitized with our laser densitometer from Molecular Dynamics, linear over the range 0-3.0 optical density units as verified by calibrated filters. Narrow O.D. ranges can be expanded to full scale (8 bit, 0-255) if required. Scanning resolution is 100 microns/pixel; the final black & white tif images are 3.3 Mbytes each for standard format and 10 Mbytes for large format.
Price: $45/scan (but included with computer comparisons).
FLUORESCENT DENSITOMETRY (2D-ES-13A)
For imaging of Sypro Ruby or Cy dyes with a Typhoon FLA 9000 biomolecular imager.
Price: $75/scan
QUANTITATIVE COOMASSIE BLUE DENSITOMETRY (2D-ES-13B)
With a GE Healthcare Image Scanner III verified to be linear from 0-2.3 OD using Melles Griot NIST-calibrated filters. Images are scanned at 84 micron resolution.
Price: $45/scan (but included with comparisons)
ELECTRONIC PHOTOS AND FILES (2D-ES-14).
The 10 MB black & white files obtained from laser densitometry are too big to email and sometimes don’t look right by eye. Our Afga Arcus II and GE Healthcare desktop scanners give excellent color images in tif format that are useful for publication and emailing (jpg format). The price includes tif and jpg image files and a letter including the methods on a CD mailed with the dried gels. The jpg images and letter are emailed to you to provide previews of 2D patterns.
Price: $26/scan
Staining
STAINING/DRYING PROCEDURES
COOMASSIE BLUE STAINING (STP-1).
Coomassie brilliant blue R250, our default protein stain, is quite sensitive (O’Farrell, P. J. Biol. Chem. 250 : 4007, 1975). The procedure, performed according to SOP L-1510, involves an alcohol/acetic acid fix, an acetic acid rehydration and an acetic acid destain. Generally, 1 µg of purified protein gives a nice spot on a 2D gel (see our internal standard), and 0.1 ug is visible. Although not as sensitive as silver staining, this method gives quantifiable results (Burgess-Cassler et. al. Clin. Chem. 35: 2297, 1989). The linear range for plots of stain density versus ng protein is broad and the method is reproducible. This stain is compatible with mass spectrometry.
Price: No charge (Default stain).
SILVER STAINING (STP-2).
Our silver stain is based on the ammoniacal silver/formaldehyde method of Oakley et. al. (Anal.Biochem.105: 361, 1980). This method involves preliminary glutaraldehyde treatment of the slab gel to fix proteins by cross linking. The pre-treatment also adds glutaraldehyde side chains to the proteins, increasing sensitivity since these groups are sites for silver deposition (Dion A, and Pomenti A Anal. Biochem. 129:390, 1983). We find this method to be about 10 times more sensitive than Coomassie blue staining, depending on the protein, even though less protein is loaded. Generally, 50 ng of purified protein gives a highly visible spot on a 2D gel (see our internal standard). However, some proteins that are detectable with Coomassie don’t stain at all with silver. Although very sensitive, this method gives semi-quantitative rather than quantitative results. The linear range for plots of stain density versus ng protein varies widely from protein to protein and also because day to day variability in stain and background intensity is observed. Silver staining is not compatible with mass spectrometry because of the glutaraldehyde.
Price: STP-2: Standard format, $63 per gel; STP-2LF: Large format, $75 per gel.
SPECIAL SILVER STAINING FOR MASS SPECTROMETRY ANALYSIS (STP-3).
This special silver stain (O’Connell and Stults Electrophoresis 18: 349-359, 1997) omits the glutaraldehyde step and is compatible with mass spectrometry fingerprinting. It often negatively stains proteins and so is not appropriate for computerized analysis. However, the pattern can be matched to our regular stain without difficulty. Note, that silver deposition interferes with the enzymatic digestion required for mass spectrometry for most mammalian proteins. If possible, use the less sensitive but more reliable Coomassie stain.
Price: STP-3: Standard format, $63/gel; STP-3LF: Large format, $75/gel.
SYPRO RUBY FLUORESCENT STAIN (STP-4).
This Molecular Probes stain fluoresces linearly with protein over a range of 100-1000 ng and is compatible with mass spectrometry. See fluorescent densitometry (2D-ES-13A) for imaging or gallery for examples.
Price: STP-4: Standard format, $118 per gel; STP-4LF: Large format, $157 each
CY DYE STAINING (STP-5).
For 2D fluorescence difference gel electrophoresis (DIGE) analysis. See DIGE, Silver, Coomassie Staining Comparison (PDF) for additional information.
Price: Inquire for quote.
STP-8: TRANSPARENCY DRYING.
All stained gels are air dried between cellophane sheets unless otherwise requested. Although this treatment adds a day to turnaround, the advantages of easier storage, little or no curling, good color retention, overlay capability, scanning capability, and overhead projection capability offset the time delay.
Price: No extra charge.
PAPER DRYING (STP-9).
This is an optional drying method for stained gels.
Price: $17/ gel
WET GELS (STP-10).
In some cases the return of wet gels is requested, for example, for a core facility’s automatic spot picker. This is no problem. The wet gels are placed between sheets of thick wet filter paper and then placed in a ziploc bag. The bag is supported by pressboard sheets and cushioned by bubble wrap prior to express mailing. Gel identification information is written on the ziploc bag.
Price: Add $35 to shipping charges
Blotting
1D and 2D GEL WESTERN BLOTTING PROCEDURES
TRANSBLOTTING ONTO PVDF (BP-1).
PVDF (Immobilon from Millipore Corp.) binds proteins more tightly than nitrocellulose and gives better recoveries. Transblotting onto PVDF is carried out overnight according to standard procedures using a CAPS/methanol buffer system without SDS. The dried stained (see BP-2) or unstained blot is returned to the client. Staining facilitates matching to a duplicate Coomassie gel for mass spectrometry. Original gels are stained to make sure they’re blank and discarded, unless otherwise requested. We will change the blotting procedure to your specifications if necessary (see 2D Gel Western Blotting for method details).
Price: BP-1: Standard format, $119 per blot; BP-1LF: Large format, $137 per blot
PVDF COOMASSIE STAINING (BP-2).
The 2D gel pattern obtained from Coomassie blue staining of a PVDF membrane is faded, but the scanned image can be contrasted with Adobe Elements. The image is super-imposable with the ECL film from Western blotting and helps to match a few spots on the film to a busy 2D gel pattern on a duplicate Coomassie gel run for mass spectrometry. Since Western blotting can be 100 times more sensitive than Coomassie, exact matching is very important. Coomassie staining of the blot does not interfere with subsequent Western blotting.
Price: add $23/blot for staining + $26/blot for electronic photos ($47/blot total)
TRANSBLOTTING ONTO NITROCELLULOSE (BP-3).
Transblotting is carried out overnight according to standard gel procedures using a tris/glycine/methanol buffer system containing 0.1% SDS, with nitrocellulose of 0.45 micron pore size. Dried unstained blots are returned to the client, and original gels are discarded unless additional treatments are requested. We will change the blotting procedure to your specifications if necessary.
Price:BP-3: Standard format, $119 per blot; BP-3LF: Large format, $137 per blot
PRE-STAINED MOLECULAR WEIGHT MARKERS FOR BLOTS (BP-4).
These markers, Precision Plus Protein Standards from BioRad, are added during slab gel electrophoresis so that colored bands are visible on both the unstained gels and subsequent transblots at molecular weights of 250, 150, 100, 75, 50, 37, 25, 20, 15, and 10 kDa.
Price: $28 per gel
WESTERN (IMMUNO-) BLOTTING, CLIENT SUPPLIES ANTIBODY (BP-6).
This method for immuno-detection of specific bands or spots is carried out on PVDF transblots from 1D or 2D gels according to standard operating procedures. Requires prior electrophoresis (2D-ES-1 or 1D-ES-1), but PVDF transblotting (BP-1) is included in the fee. Generally the client supplies the antibody. The sensitive ECL method of detection is used.
Price: BP-6SF: Standard format, Add $500/sample
BP-6SF-Dup: Duplicate SF blot, Add $343
BP-6LFLarge format, Add $623/sample
BP-6LF-Dup: Duplicate LF blot, Add $474
PHOSPHOTYROSINE WESTERN BLOTTING PACKAGE INCLUDING PY20 ANTIBODY (BP-7). Phosphotyrosine Western blotting
P Tyr Western blotting has proven to be remarkably sensitive, specific and reproducible. Package includes 2D gel electrophoresis, transblotting to PVDF, PVDF Coomassie staining, immunostaining with PY20 antibody, and electronic photos of stained PVDF and ECL films.
Prices: BP-7SF: Standard Format, $592/sample
BP-7SF-Dup: Duplicate SF package, $406 each
BP-7LF: Large format,$672/sample
BP-7LF-Dup: Duplicate LF package, $500 each
COMBINED P-SERINE/P-THREONINE 2D WESTERN BLOTTING (BP-8).
Package includes ECL ultra and duplicate blots but not antibodies. We have optimized a method for 2D Western blotting against phosphoserine and phosphothreonine residues on proteins using the Q5 (P-Serine) and Q7 (P-Threonine) antibodies from Qiagen and ECL Ultra from Lumigen. Both antibodies are reported to detect phosphorylated residues irrespective of surrounding amino acids. A control sample should be run for purposes of comparison. The high sensitivity adds variability, so all BP-8 packages include duplicate Western blots to verify results. Antibodies (not included) are purchased fresh (at cost) for each client. Package includes duplicate 2D gels with PVDF transblots, PVDF Coomassie staining, immunostaining with combined Q5/Q7 antibodies, and electronic photos of stained PVDFs and ECL films.
Price: Inquire to discuss project & receive a quote.
PHOSPHOSERINE OR PHOSPHOTHREONINE WESTERN BLOTTING PACKAGE (BP-9).
Same package as BP-8 except a single antibody. Duplicate blots included.
Price: Inquire to discuss project & receive a quote.
ACETYL-LYSINE WESTERN BLOTTING PACKAGE (BP-10).
Lysine acetylation is an important post translational modification occurring on many proteins. Our 2DE Acetyl-lysine Western blot package allows you to identify acetylated proteins in a complex mixture using polyclonal antibody 9441 from Cell Signaling Technology. Package includes 2DE, transblotting to PVDF, PVDF Coomassie staining, subsequent immunostaining with anti-acetyl-lysine antibody, and electronic photos of stained blot and ECL films.
Price: BP-10SF: Standard format, $721/each;
BP-10SF-Dup: SF Duplicate package, $577/sample
BP-10LF: Large format, $922/sample;
BP-10LF-Dup: LF Duplicate package, $721 each
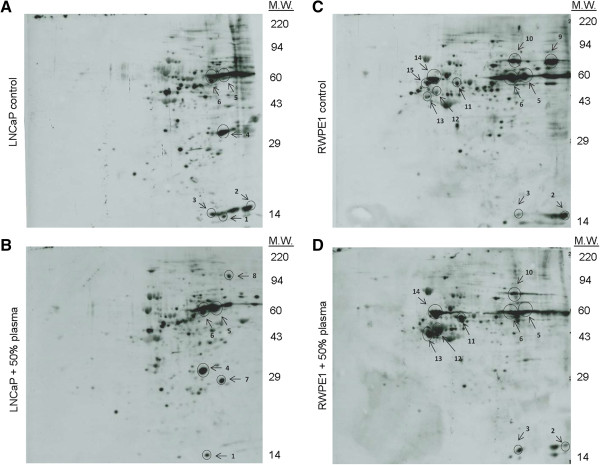
Image taken from: Mitra, R., Goodman, O., and Le, T. Enhanced detection of metastatic prostate cancer cells in human plasma with lipid bodies staining. BMC Cancer, 2014. 14: p. 91. Figure 2. 2D Western blots of protein lysine acetylation profiles. Lysine acetylation profiles of (A) untreated LNCaP cells, (B) LNCaP cells incubated with human plasma, (C) untreated RWPE1 cells, and (D) RWPE1 cells incubated with human plasma.
HCP
HCP Antibody or Drug Substance Analysis using 2D Electrophoresis
For additional information see HCP overview, HCP method, HCP results/report, and product testing or the PowerPoints:2DE and HCP Antibody Analysis, Silver-filmAlignment, and HCP Method provide more information about uses and methods of using 2DE for HCP analysis.
ELISA Antibody Reactivity with Complex Null Cell or DS-Containing Whole Cell Lysates
These tests are time-consuming because of 2D pattern complexity (often >1000 protein spots) for both silver and Western blot images. Any protein spots unique to the Western blot images are shown as blue dots on the silver master pattern in the report. Each 2DE package includes optimized Western blot with light and dark exposures, silver-stained 2D gels in duplicate, electronic photos, and computer comparison with complete report including percent coverage. HCP Example Report (PDF).
Price: HCP-WCL-SF:(Standard Format) $4146 includes complete analysis of one HCP antibody using standard format 2D gels (13 x 15 cm)
HCP-WCL-SFA: $3373/Additional antibody, same SF silvers
HCP-WCL-LF: (Large Format) $4609 includes complete analysis of one HCP antibody using large format 2D gels (20 x 22 cm)
HCP-WCL-LFA: $3739/Additional antibody, same LF silvers
Semi-Purified or Final Drug Substance Analysis Packages
These packages, similar to those for the ELISA antibody analyses, are useful for confirming product purity. Note that DS breakdown products are removed from the pattern by comparison to an additional anti-DS Western blot (HCP-PSR-LFR below), if necessary. Each 2DE package includes an optimized Western blot with light and dark exposures, silver-stained 2D gels in duplicate, electronic photos, and computer comparison with complete report. HCP Example Report (PDF)
Price: HCP-DS-SF:(Standard Format) $3049 includes complete analysis of one HCP antibody using standard format 2D gels (13 x 15 cm)
HCP-DS-SFA:$2441/Additional antibody, same SF silvers
HCP-DS-LF: (Large Format) $3368 includes complete analysis of one HCP antibody using large format 2D gels (20 x 22 cm)
HCP-DS-LFA: $2745/Additional antibody, same LF silvers
Product Spot Removal (HCP-PSR-LFR)
Removal of product spots from HCP-LF report. Includes an additional Western blot obtained with anti-product antibody.
Price: Add $1298
HCP-PSR-LFRA: Removal of product spots for additional HCP antibody using product Western blot obtained with anti-product antibody.
Price: Add $644
Run Documentation (GTRK-1) Add 30% for documentation/buffer tracking as provided on IEF, slab gel, and Western blot checksheets (PDF) for HCP-related analyses. Organized archival of reports and data files on the password protected cloud storage site Egnyte is included with run documentation. We’ll send you an invitation to the link; you then create a new password for your data.
Price: Add 30%
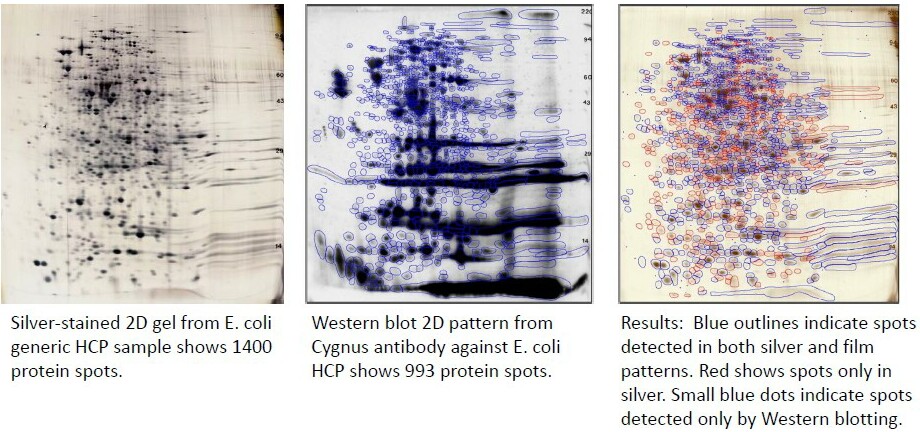
Example of Images from an E. coli HCP Antibody Analysis; 993/1400 total spots are detected by the antibody = 71% coverage.
HCP Antibody Analysis using 1D Electrophoresis (SDS PAGE)
Anti-HCP antibodies are polyclonal, usually from rabbits or goats, because the antigenic protein mixture is complex. 1D-HCP is a good way to check the time course of HCP antibody production and to compare immunogenic reactions of different animals.
1D-HCP-1. 1D package includes: HCP sample preparation, standard format SDS PAGE (13 x 15 cm gels, 15 wells), Western blot with light and dark exposures, Coomassie blue-stained gel, electronic images, and a printed report showing images. Western blot optimization is free. Visual interpretation, usually straightforward, is performed by the client unless otherwise requested.
Price: $773 each
1D-HCP1A: Additional antibodies tested against the same antigen.
Price: $548 each
Sample Prep
Sample Preparation Procedures
General Comments
-SDS Buffer works well. Carrier ampholine IEF in tube gels is compatible with 2.5% SDS. See SDS Compatibility explanation.
-1D Samples work well. Any sample that you’ve previously prepared for SDS PAGE gels can usually be run on our 2D gels as long as the protein concentration is known and salt is < 200 mM. To make sure, send an image of the 1D gel to Jon Johansen, Lab Manager (jon@kendricklabs.com), with lanes identified. Send samples frozen on dry ice with a sample identification form.
-IP Bead Prep is free. We will take proteins off your immunoprecipitation beads without charge prior to running 2D gels. Just take the supernatant off and send the beads on dry ice with a sample ID form.
-Advice is free. Don’t hesitate to call for advice if you’d like to prepare samples in your lab to save money. Buffer recipes are posted on our web site, or order our Mailing Kit (MK-1, $380 includes dry ice shipping) for premade buffers. If there’s a problem with sample preparation, we’ll do free reruns to optimize the pattern.
Services
PROTEIN DETERMINATION (SP-1):
The total amount of protein in each sample is determined by the Pierce BCA total protein assay (Smith, P., et. al., Anal Biochem, 1985. 150:76-85). The BCA assay is compatible with SDS but not reducing agents. If you require protein determinations, do not dissolve samples in buffers containing dithiothreitol (DTT) or beta-mercaptoethanol (BME). We will add BME after protein analysis.
Samples containing dithiothreitol are incompatible with the BCA assay but may be analyzed with a Coomassie dye binding assay (Bio-Rad Labs; Bradford, M., Anal Biochem, 1976. 72 :248-54). Because this assay is incompatible with SDS, proteins are often not dissolved well. The BCA test is preferred, if possible.
Price: $62 per sample (BCA or Bradford).
COMBINING MULTIPLE SAMPLE TUBES (SP-2).
For example, from an immunoprecipitation.This involves transferring each tube’s contents, adding a rinse, vortexing, and transferring the rinse. The rinsing step is repeated as needed.
Price: 1-4 tubes, no charge; 5-10 tubes, $75/sample; 11-20 tubes, $149/sample.
ETHANOL PRECIPITATION OF PROTEINS (SP-3): Precipitation is followed by buffer addition. See the Ethanol Precipitation (PDF) for more information.
Price: $40 per sample.
HOMOGENIZATION (SP-4).
Includes homogenization with osmotic lysis buffer containing protease inhibitors followed by incubation on ice with DNase and RNase to break down interfering polynucleotides. The composition of the 10X Nuclease Stock Solution is given in our Sample Preparation Guide. If you require nuclease treatment, send the samples in a buffer containing less than 0.3% SDS. Higher SDS concentrations inactivate nucleases.
Price: $57 per sample.
MICRODIALYSIS (SP-5)
Followed by lyophilization & buffer addition. High salt concentration (> 150 mM) interferes with IEF; the less salt the better. If your samples contain high salt or buffer (phosphates, NaCl, etc.), this dialysis and concentration step is highly recommended.
Price: $45 per sample.
LYOPHILIZATION (SP-6):
Followed by buffer addition.
Price: $40 per sample.
TCA PRECIPITATION (SP-7):
Followed by buffer addition.
Price: $40 per sample.
TCA/ACETONE PRECIPITATION (SP-8):
Followed by buffer addition.
Price: $40 per sample.
CUSTOM SAMPLE PREPARATION (SP-9):
(Chunks of tissue, leaves, etc.) Charged by the hour.
Price: $249/hr.
ALBUMIN/IgG REMOVAL FROM SERUM, PLASMA & CSF OF HUMAN, RAT AND MOUSE (SP-10):
We use the Millipore Sigma ProteoExtract Albumin/IgG Removal kit (Cat. No. 122642). Depletion of albumin and IgG from 20-60 ul using the disposable gravity-flow affinity columns removes up to 75% of total serum proteins so that 3-4 times more enriched sample can be loaded on 2D gels.
Price: $95/sample.
BioRad Ready Prep (SP-11):
Price: $117/sample.
Mass Spectrometry
Protein Identification by Mass Spectrometry
We have considerable experience in sending 2D gel proteins to university and commercial core facilities for identification by mass spectrometry, LC-MS/MS. Turnaround time for mass spectrometry is usually about two weeks. We highly recommend Clarkson University Proteomics Laboratory. Contact Dr. Costel Darie, the Facility Director, for more information (call 315-268-7763 or email:cdarie@clarkson.edu).
SPOT CUTOUT FOR MASS SPECTROMETRY (2D-ES-19):
We do this manually and are obsessively careful. The cutout spots are placed in Eppendorf tubes for express mail to the core facility of your choice; our default is the Clarkson University Proteomics Lab. The cost includes a letter listing the spots to the core facility, a copy of the letter for you, and an image or original gel showing spot locations. Each spot is given a unique identifier.
Price: $35 per spot
MASS SPECTROMETRY using LC-MS/MS (2D-ES-18):
This powerful method is used to identify proteins or to determine the unknown amino acid sequence of peptide fragments of the protein of interest cut from Coomassie blue or special silver-stained gels. This method is especially recommended for spots or bands which might contain mixtures and for samples from species with incomplete genomes. Protein spots are cut from Coomassie blue or special silver-stained gels and digested with trypsin. The digestion mixture is separated on an LC Packings nano-lc on which the detector outlet is connected directly to the nanospray source of a Micromass Q-Tof mass spectrometer. Peptides are eluted at a flow rate of 200 nl/min and are scanned as they enter the source. When a peptide is detected, the Q-Tof is programmed to switch to MS/MS mode, which means that the eluting peptide is fragmented by means of an applied collision energy, and the resulting ions are scanned for several seconds. When the programmed time for MS/MS is done, the Q-Tof switches back to MS mode and resumes scanning the eluting peptides. Because peptides fragment at the peptide bond in a predictable way, the fragmentation pattern can be used to deduce sequence information. The ions resulting from fragmentation along with the mass of the intact peptide can be used to search a database and identify the protein.
Price: 2D-ES-18 LC-MS/MS: 1-10: $305 per polypeptide spot
2D-ES-18a LC-MS/MS: 11 or more: $268 per polypeptide spot
Reference for protein identification by LC-MS/MS
1. Hernandez, P, Muller, M, and Appel, RD. Automated protein identification by tandem mass spectrometry: issues and strategies. Mass Spectrom Rev. 25: 235-254 (2005).
IDENTIFICATION OF POST-TRANSLATIONAL MODIFICATIONS (PTM) USING BIOINFORMATICS SOFTWARE (2D-ES-18-PTM):
Once proteins have been identified by LC-MS/MS, analysis of PTMs may be performed using bioinformatics software. PTMs that can be targeted include methylation, acetylation, phosphorylation, sulfonylation, glycosylation, ubiquitination, sumoylation, and disulfide bridges. In most experiments, the protein investigated has two states for a PTM site: unmodified and modified. Data analysis is performed with a specially constructed database, which will look for PTMs within the protein using unbiased searches (100% chance for an amino acid to be both modified and unmodified). If the amino acid is modified, the modified peptide will be identified, but not the unmodified one and vice versa. Assuming the peptides fly, PTM’s can usually be identified. Other mass spectrometry methods especially designed for PTM analysis, such as product ion discovery or neutral loss, can also be performed. Includes analysis of LC-MS/MS spectra for up to 3 PTMs. Requires prior 2D-ES-18 for both modified and unmodified samples. Please call Dr. Darie (315-268-7763) to discuss PTM analysis before sending samples.
Price:2D-ES-18-PTM: $731 per sample
CUSTOM INTERPRETATION (2D-ES-20):
Consulting fee for the interpretation of of complex spectra by Clarkson Personnel.
Price:2D-ES-20: $249/hr
Misc
Miscellaneous Services
CURRENT GOOD MANUFACTURING PRACTICE (CGMP) :
The numerous SOPs required for CGMP for 1D and 2D electrophoresis are in place at Kendrick Labs along with an archival system for equipment calibrations and reports (see Accreditation). However, custom validation for the sample type of interest including SOPs are required for new projects to pass detailed FDA inspections – which may occur years down the road.Please let us know if you are going to submit an IND or plan to use our lab for QA of a product.
Price: Request a quote, varies with project.
CONSULTING FEES (CON-1)
Price: $249/hr.
ARCHIVE-1:
Archival of reports and data files for 3 years on our password protected cloud website, Egnyte. Note that this is included in item buffer tracking (GTRK-1) for HCP antibody analyses.
Price: $110/3 years
COMING SOON: A new phosphotyrosine internal standard
Price: Call/email for more information